Graduation project at the Visualization Group of Eindhoven University of Technology, with the goal add genome size data support to the existing DNAVis2 sequence browser. The basic DNAVis2 sequence browser is an OpenGL accelerated visualization tool to visualize and explore a small number of annotated DNA sequences. The application is written in Java using the NetBeans framework and the JOGL OpenGL bindings.
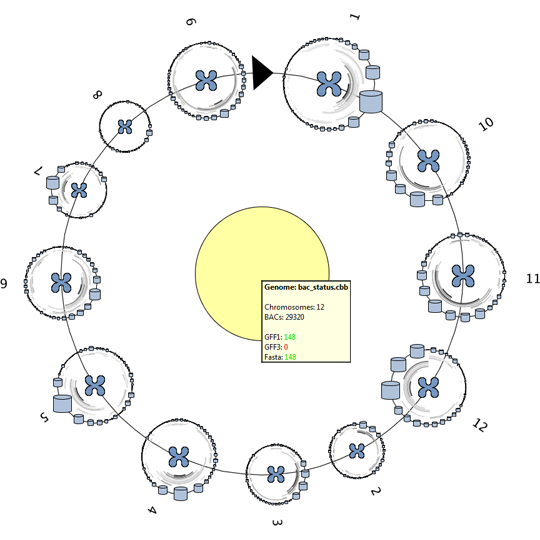
The extended version is designed and implemented during this master project and adds the data structures and visualizations needed to visualize and explore the data of a complete genome, consisting of 10.000-100.000 annotated DNA sequences. Two approaches are used to provide more inside into the dataset as a whole.
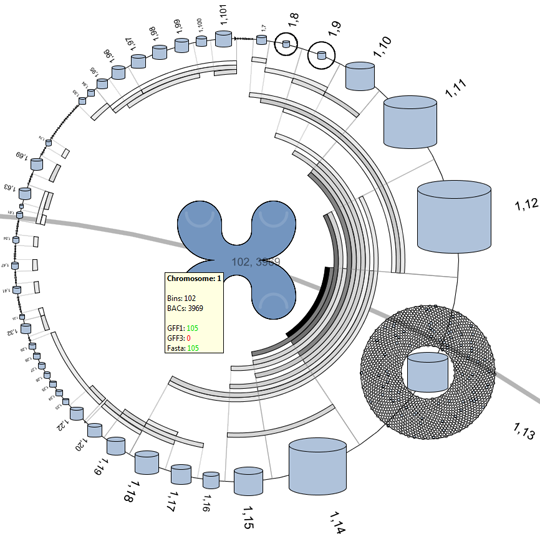
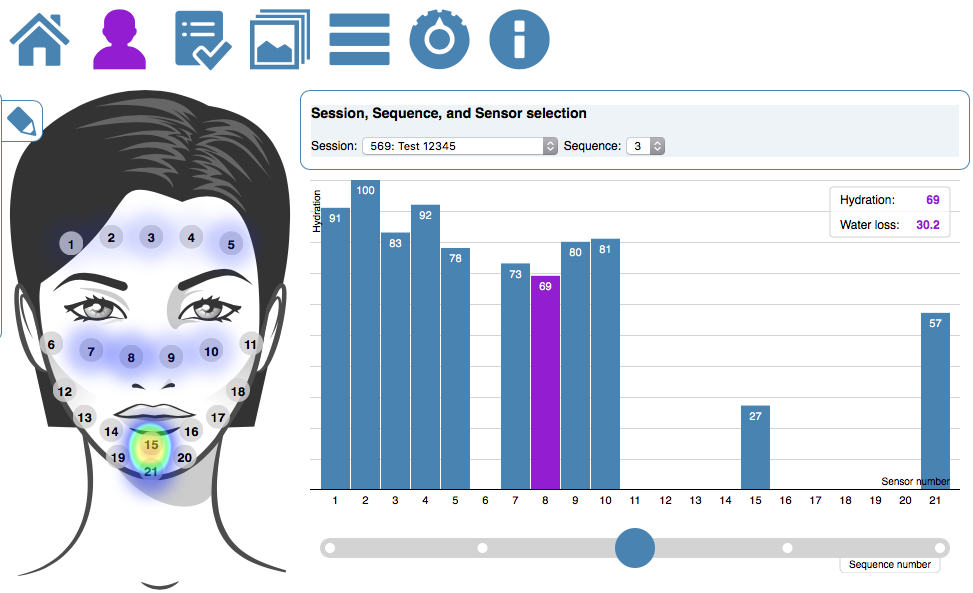
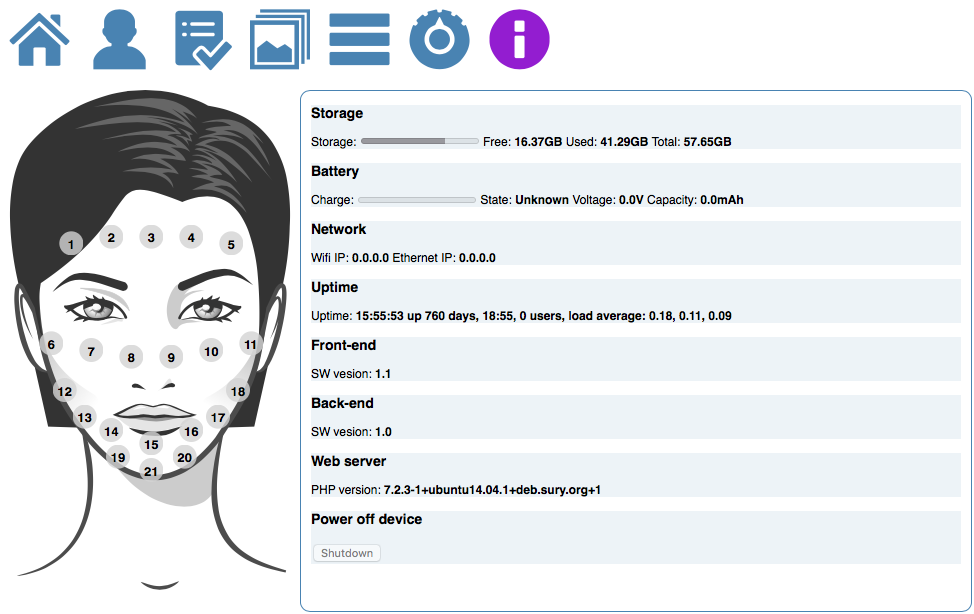
The first approach makes use of the higher level DNA structures to visualize the data distribution and to provide a high level interface to lower level annotations and sequencing data. This provides the biologist with the tools to browse and create sub-selection using the DNA’s structural properties.
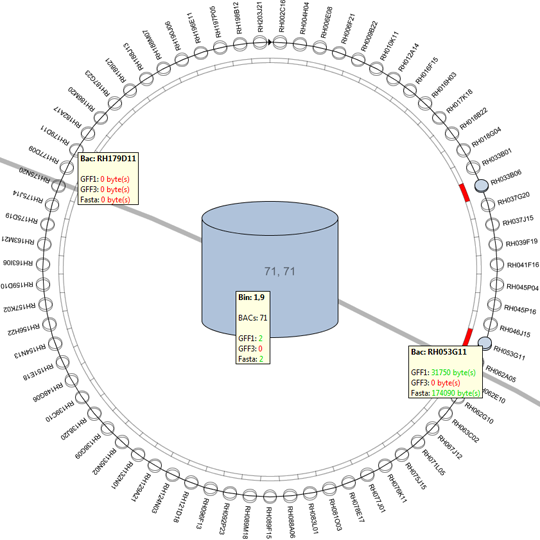
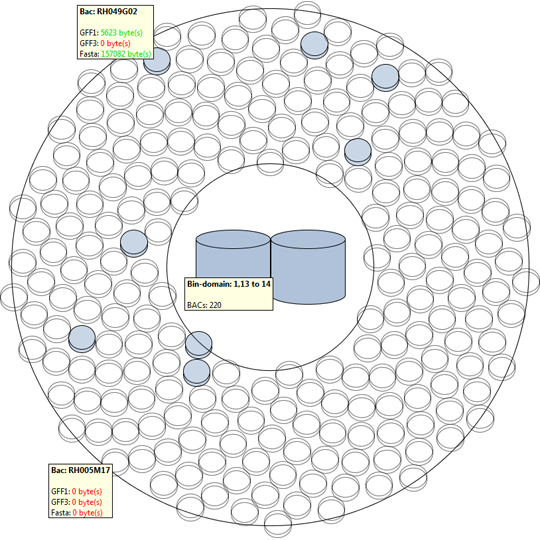
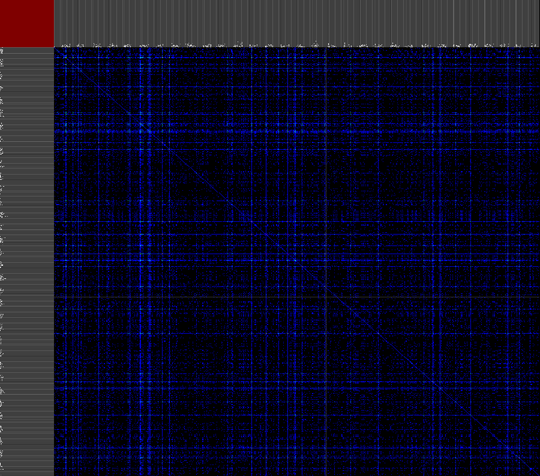
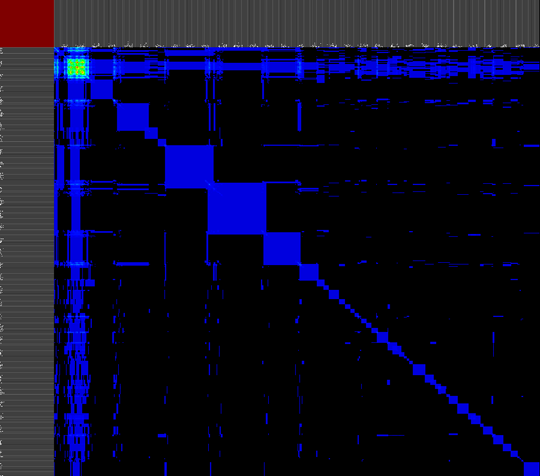
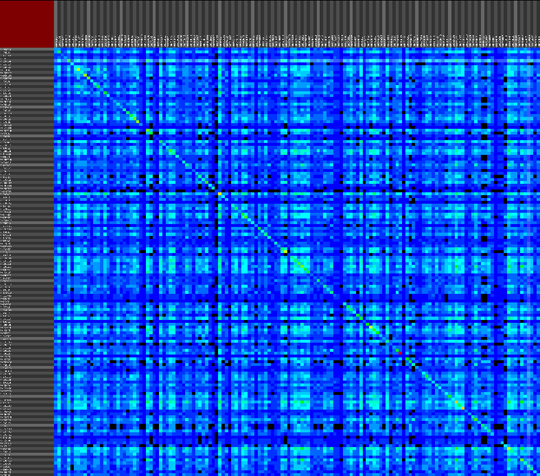
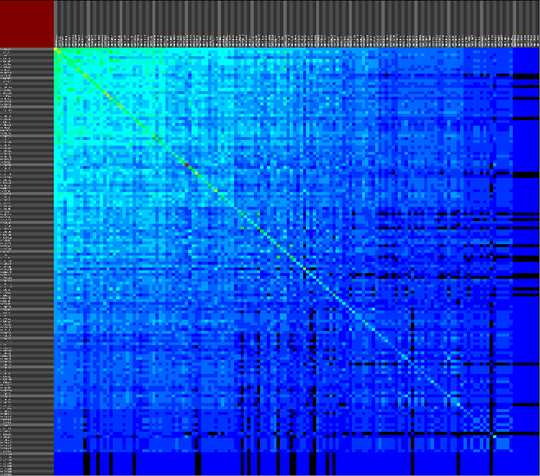
The second approach provides a versatile interface to cross-reference various data properties across all abstraction levels. This tool provides the biologist with an inside into the annotation data of the dataset as a whole (or a predefined sub-selection of the data), this results cross-reference visualizations than can contain more then 10.000 row and column items.